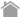Protein-Protein Interaction Detection System "CoralHue™ Fluo-chase Kit"
Protein-Protein Interaction Detection System
- Protein-protein interaction (PPI) analysis
- Direct visualization of PPIs in living cells
- Easy construction
CoralHue™ Fluo-chase Kit is based on the protein fragment complementation assay using our proprietary fluorescent protein , CoralHue™ monomeric Kusabira Green (mKG).
mKG is split into two inactive fragments, mKG_N and mKG_C, and do not emit
fluorescence on their own.
With the CoralHue™ Fluo-chase Kit the protein of interest (A) is expressed as a fusion protein
to the mKG_N (A-mKG_N) and the mKG_C is fused to the second protein of interest (B),
(B-mKG_C).
Both vectors encoding mKG fragment fusion protein are co-introduced into mammalian cells. When the mKG fragments are in close proximity due to the interaction of A and B, the mKG fragments form a beta-barrel structure and emit green fluorescence.
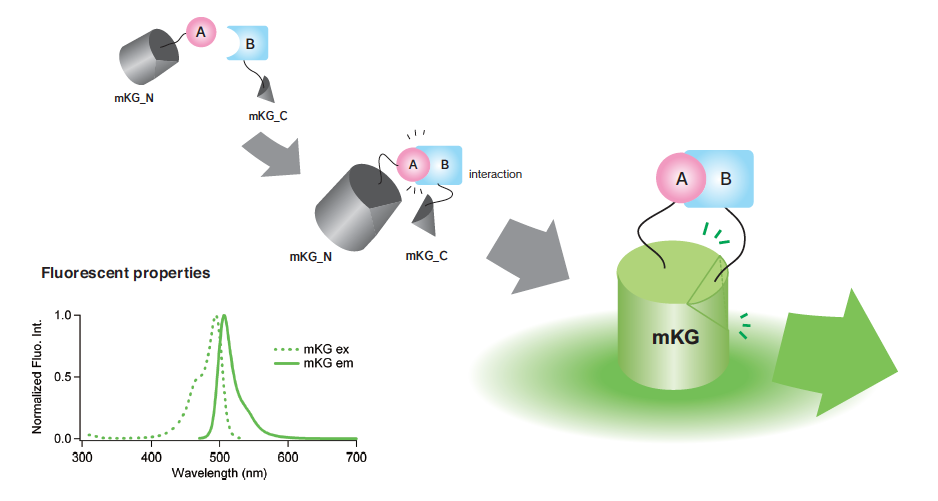
mKG_N and mKG_C (left panel) or p65-mKG_N and p50-mKG_C (right panel) were co-expressed in HEK293T cells. After a 24h incubation, cells were imaged on an inverted microscope. HEK293T cells expressing the N terminal fragment of mKG (mKG_N) and/or the C terminal fragment of mKG (mKG_C) absolutely had no fluorescent signal.(p65-mKG_N ; 190 to 291 amino acid sequence of p65(RELA) fused to mKG_N. p50-mKG_C ; 247 to 352 amino acid sequence of p50(NFKB1) fused to mKG_C.)
There is the possibility of a nonspecific mKG fluorescent signal from an unexpected protein-protein interaction. For example, in our original experiments, HEK293T cells expressing both the mKG_N with a flexible linker and the p65-mKG_C emitted a nonspecific fluorescent signal. The intensity of the signal was about 20% of the p65-mKG_N and p50-mKG_C co-expressing HEK293T cells. Why the unexpected interaction occurred is now under evaluation.
| Open reading frame (bp) | Molecular weight (kDa) | |
|---|---|---|
| mKG_N | 510
|
19.0
|
| mKG_C | 156
|
5.7
|
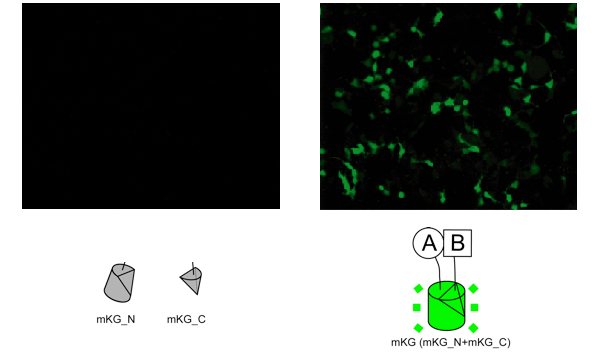
For a topological variety of mKG fragments and proteins of interest, CoralHue™ Fluo-chase Kit contains four regioselective vectors.Using the vector sets, all eight possible patterns for the proteins of interest (A and B) can be examined to select the most favorable fusion construct.
The lower panel shows all possible patterns between the cell cycle-related factors of p25 and Cdk5 interaction. Two of the eight pairs (mKG_N-Cdk5 + mKG_C-p25 and mKG_N-Cdk5 + p25-mKG_C ) successfully emitted bright fluorescent signals however, two other pairs (mKG_C-Cdk5 + mKG_N-p25 and mKG_C-Cdk5 + p25-mKG_N ) failed to emit a fluorescent signal.
Detection of PPIs using CoralHue™ Fluo-chase Kit largely depends on the property of the protein complex, such as molecular mass, the direction of the N and C terminals, molecular proximity to each terminal, linker sequence, and so on. This product does not ensure the ability to detect all PPIs in nature.
| Excitation/emission maxima |
Molar extinction coefficient (M-1cm-1) |
Fluorescence quant yield |
pH sensitivity in fluorescence |
Number of amino acid |
|
|---|---|---|---|---|---|
| mKG | 494/506
|
64,200 (494 nm)
|
0.57
|
pKa=6.1
|
218
|
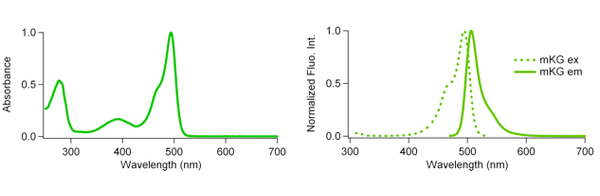
| monomeric Kusabira-Green(mKG) spectle data files (txt files) | mKG ex.( |
| mKG em. ( |
| Code No. | Products | Vol. |
|---|---|---|
| AM-1100M | CoralHue™ Fluo-chase Kit | 1 system |
| Components |
|
|
| CHARACTERISTIC | Kusabira-Green |
|---|---|
| Fluorescence color | Green |
| Oligomerization | monomer |
| Excitation max (nm) | 494 |
| Emission max (nm) | 506 |
| Molar extinction coefficient (M-1cm-1) | 63,200 (494 nm) |
| Fluorescence quantum yield | 0.57 |
| Brightness*1 | 36.0 |
| pH sensitivity | pKa=6.1 |
| Molecular weight (kDa) | 24.5 |
*1Brightness: Molar extinction coefficient x Fluorescence quantum yield /1000.
CoralHue™ mKG_N and mKG_C can be recognized using antibodies as shown below.
- Anti-monomeric Kusabira-Green N-terminal fragment, Code No. M148-3M
- Anti-monomeric Kusabira-Green C-terminal fragment, Code No. M149-3M
※CoralHue™ Kusabira-Green was co-developed with the Laboratory for Cell Function and Dynamics, the Advanced Technology Development Center, the Brain Science Institute, RIKEN.
※Improvement of monomeric Kusabira-Green is an accomplishment of co-development by NEDO project (Project code: P06008, Team leader: Dr. Natsume Tohru)
※If you belong to a profit organization or a business corporation, or wish to use the product for profit or commercial purposes, please contact us.
[Marketing Department RUO Group]
TEL: +81-3-6854-3614 E-mail: support@mbl.co.jp
References
- Ueyama T, Kusakabe T, Karasawa S, Kawasaki T, Shimizu A, Son J, Leto TL, Miyawaki A, Saito N. Sequential Binding of Cytosolic Phox Complex to Phagosomes through Regulated Adaptor Proteins: Evaluation Using the Novel Monomeric Kusabira-Green System and Live Imaging of Phagocytosis. J Immunol. 181: 629-640 (2008) PMID: 18566430
- Hamatake M, Aoki T, Futahashi Y, Urano E, Yamamoto N, Komano J. Ligand-independent higher-order multimerization of CXCR4, a G-protein-coupled chemokine receptor involved in targeted metastasis. Cancer Sci. 100: 95-102 (2009) PMID: 19018754
- Hashimoto J, Watanabe T, Seki T, Karasawa S, Izumikawa M, Seki T, Iemura S, Natsume T, Nomura N, Goshima N, Miyawaki A, Takagi M, Shin-Ya K. Novel in vitro protein fragment complementation assay applicable to high-throughput screening in a 1536-well format. J Biomol Screen. 14: 970-979 (2009) PMID: 19641222
- Yoshida T, Ebina H, Koyanagi Y. N-linked glycan-dependent interaction of CD63 with CXCR4 at the Golgi
apparatus induces downregulation of CXCR4. Microbiol Immunol. 53: 629-635 (2009) PMID: 19903263
※CoralHue™ fluorescent proteins were co-developed with the Laboratory for Cell Function and Dynamics, the Advanced Technology Development Center, the Brain Science Institute, RIKEN. MBL possesses the license and deals in the products.
Please send us a completed and signed License Acknowledgement.
Please note that the product will be shipped after we receive a
completed and signed License Acknowledgement (PDF file on the right).
Prior to shipping, our staff may contact you to verify the intended use
of the product. Please also note that a separate agreement or a contract
may be required depending on the intended use.
The product you have ordered is sold for research purpose only. It is
not intended for industrial or clinical use, and shall not be used for
any purposes other than research. You are also asked not to hand over or
resell the product to a third party.
If you belong to a profit organization or a business corporation, or
wish to use the product for profit or commercial purposes, please
contact us.

- Fluorescent protein
- Basic fluorescent proteins
- Midoriishi-Cyan [Outline] [Product list]
- Umikinoko-Green [Outline] [Product list]
- Azami-Green [Outline] [Product list]
- Kusabira-Orange [Outline] [Product list]
- dKeima570 [Outline] [Product list]
- Keima-Red [Outline] [Detection of mitophagy with Keima-Red] [Product list]
- Photoconvertible fluorescent proteins
- Dronpa-Green [Outline] [Product list]
- Kaede [Outline] [Product list]
- Kikume Green-Red [Outline] [Product list]
- Organelle targeting vectors [Product list]
- Advanced fluorescent indicators
- Image based Protein-Protein interaction analysis "Fluoppi" [Outline] [Fluoppi Red] [Product list]
- Cell cycle indicator "Fucci" [Outline] [Product list]
- Protein-Protein Interaction Detection System "CoralHue™ Fluo-chase Kit" [Outline] [Product list]
- Anti-CoralHue™ fluorescent protein antibodies
- Anti-CoralHue™ fluorescent protein antibodies [Outline] [Product list]
- Resources
- CoralHue™ vectors full DNA sequence
- Spectrum