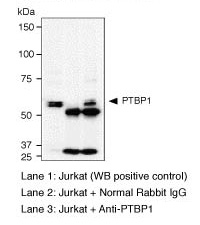
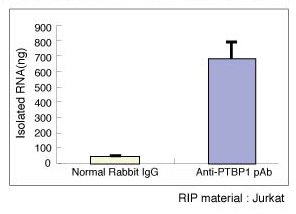
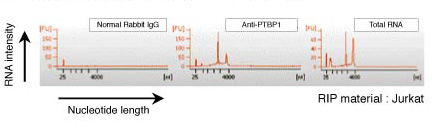
| Notes |
In the RIP-Assay protocol, mRNP complexes are isolated from cell extracts by immunoprecipitation with RIP-Certified Anti-RBP Antibodies provided from MBL. mRNAs are isolated from mRNPs using guanidine hydrochloride. Thus, RIP-Assay Kit does not contain phenol or chloroform, allowing safe isolation of “high-quality RNA” from RNP complexes without degradation. Once purified, the RNAs present in the complex are analyzed to identify the target mRNAs using various molecular biology tools such as RT-PCR, gene expression analysis based on microarray technology (Chip analysis), or sequencing.
The major advantage of RIP-Assay Kit over most other omics approaches is that the majority of the RNAs identified exhibit structural and functional relationships. Structurally, the mRNAs in a complex contain common binding motifs for the RNA binding protein employed. Functionally, the RNAs identified generally share a common RNA regulatory network as they tend to be co-localized by the RNA binding protein which may determine how they are utilized in the cell.
A second advantage of RIP-Assay Kit over traditional RNA isolation and analysis methods is that the fractionation procedure effectively concentrates the RNA species bound to a specific binding protein enabling small changes in levels of low-abundance RNAs to be detected with a greatly increased signal-to-noise ratio.
When performed according to the supplied protocol, another advantage of RIP-Assay Kit is that RNA reassociation is minimized and RNAs contained in the RNP complex of interest are abundantly recovered. |
| Citations |
- Thompson K et al. Acute adaptive responses of central sensorimotor neurons after spinal cord injury. Translational Neuroscience 1, 268-78 (2010)
- Jin D et al. RNA-binding motif protein 24 regulates myogenin expression and promotes myogenic differentiation. Genes Cells 15, 1158-67 (2010)(PMID:20977548)
- Lachke SA et al. Mutations in the RNA granule component TDRD7 cause cataract and glaucoma. Science 331, 1571-6 (2011)(PMID:21436445)
- Nouvade R et al. Activation of p38 MAPK in CD4 T cells controls IL-17 production and autoimmune encephalomyelitis. Blood 118, 3290-3300 (2011)(PMID:21791428)
- Ohno S et al. Polypyrimidine tract-binding protein regulates the cell cycle through IRES-dependent translation of CDK11(p58) in mouse embryonic stem cells. Cell Cycle 10, 3706-13 (2011)(PMID:22037210)
- Hayman TJ et al. Translation initiation factor eIF4E is a target for tumor cell radiosensitization. Cancer Res. 72, 2362-72 (2012)(PMID:22397984)
- Miyazaki Y et al. Viral delivery of miR-196a ameliorates the SBMA phenotype via the silencing of CELF2. Nat Med. 18, 1136-41 (2012)(PMID:22660636)
- Janiszewska M et al. Imp2 controls oxidative phosphorylation and is crucial for preserving glioblastoma cancer stem cells. Genes Dev. 26, 1926-44 (2012)(PMID:22899010)
- Matsui T et al. Celf1 regulation of dmrt2a is required for somite symmetry and left-right patterning during zebrafish development. Development 139, 3553-60 (2012)(PMID:22899848)
- Xu L et al. A novel function of RNAs arising from the long terminal repeat of human endogenous retrovirus 9 in cell cycle arrest. J Virol. 87, 25-36 (2013)(PMID:23097441)
- Zhang LY et al. MicroRNA-144 promotes cell proliferation, migration and invasion in nasopharyngeal carcinoma through repression of PTEN. Carcinogenesis. 34, 454-63 (2013)(PMID:23125220)
- Banadakoppa M et al. Role of transcription factor Sp1 and RNA binding protein HuR in the downregulation of Dr(+) Escherichia coli receptor protein decay accelerating factor (DAF or CD55) by nitric oxide. FEBS J. 280, 840-54 (2013)(PMID:23176121)
- Morita M et al. The Lipid Mediator Protectin D1 Inhibits Influenza Virus Replication and Improves Severe Influenza. Cell 153, 112-25 (2013)(PMID:23477864)
- Takagi S et al. RNP2 of RNA Recognition Motif 1 Plays a Central Role in the Aberrant Modification of TDP-43. PLoS One 8, e66966 (2013)(PMID:23840565)
- Udan-Johns M et al. Prion-like nuclear aggregation of TDP-43 during heat shock is regulated by HSP40/70 chaperones. Hum Mol Genet. 23, 157-70 (2014)(PMID:23962724)
- Tahara N et al. Celf1 is required for formation of endoderm-derived organs in zebrafish. Int J Mol Sci. 14, 18009-23 (2013)(PMID:24005864)
- Sasaki-Osugi K et al. Nuclear ALG-2 Interacts with CHERP Ca2+-Dependently and Participates in Regulation of Alternative Splicing of Inositol Trisphosphate Receptor Type 1 (IP3R1) Pre-mRNA. J Biol Chem. 288, 33361-75 (2013)(PMID:24078636)
- He T et al. MicroRNA-542-3p inhibits tumour angiogenesis by targeting Angiopoietin-2. J Pathol. 232, 499-508 (2014)(PMID:24403060)
- Mizutani T et al. 7SK small nuclear ribonucleoprotein complex is recruited to the HIV-1 promoter via short viral transcripts. FEBS Lett. 588, 1630-6 (2014)(PMID:24607481)
- Yoo JS et al. DHX36 Enhances RIG-I Signaling by Facilitating PKR-Mediated Antiviral Stress Granule Formation. PLoS Pathog. 10, e1004012 (2014)(PMID:24651521)
- Ferrarese R et al. Lineage-specific splicing of a brain-enriched alternative exon promotes glioblastoma progression. J Clin Invest. 124, 2861-76 (2014)(PMID:24865424)
- Shi JX et al. CNOT7/hCAF1 is involved in ICAM-1 and IL-8 regulation by tristetraprolin. Cell Signal. 26, 2390-6 (2014)(PMID:25038453)
- Zhang C et al. Hepsin inhibits CDK11p58 IRES activity by suppressing unr expression and eIF-2α phosphorylation in prostate cancer. Cell Signal. 27, 789-97 (2015)(PMID:25576733)
- Chung BM et al. Regulation of C-X-C chemokine gene expression by keratin 17 and hnRNP K in skin tumor keratinocytes. J Cell Biol. 208, 613-27 (2015)(PMID:25713416)
- Kozaki T et al. Role of zinc-finger anti-viral protein in host defense against Sindbis virus. Int Immunol. 27, 357-64 (2015)(PMID:25758257)
- Somasekharan SP et al. YB-1 regulates stress granule formation and tumor progression by translationally activating G3BP1. J Cell Biol. 208, 913-29 (2015)(PMID:25800057)
- Samanta S et al. IMP3 promotes stem-like properties in triple-negative breast cancer by regulating SLUG. Oncogene 35, 1111-21 (2016)(PMID:25982283)
- Tsujumura K et al. miR-199a Links MeCP2 with mTOR Signaling and Its Dysregulation Leads to Rett Syndrome Phenotypes. Cell Rep. 12, 1887-901 (2015)(PMID:26344767)
- Jiang H et al. microRNA-185 modulates low density lipoprotein receptor expression as a key posttranscriptional regulator. Atherosclerosis. 243, 523-32 (2015)(PMID:26523989)
- Yabe-Wada T et al. TLR signals posttranscriptionally regulate the cytokine trafficking mediator sortilin. Sci. Rep. 6, 26566 (2016)(PMID:27220277)
- Ito J et al. RBM20 and RBM24 cooperatively promote the expression of short enh splice variants.FEBS Lett. 590, 2262-74 (2016)(PMID:27289039)
- Olejniczak SH et al. Coordinated Regulation of Cap-Dependent Translation and MicroRNA Function by Convergent Signaling Pathways. Mol Cell Biol. 36, 2360-7 (2016)(PMID:27354062)
- Cui J et al. PTBP1 modulation of MCL1 expression regulates cellular apoptosis induced by antitubulin chemotherapeutics. Cell Death Differ. 23, 1681-90 (2016)(PMID:27367564)
- Popovitchenko T et al. The RNA binding protein HuR determines the differential translation of autism-associated FoxP subfamily members in the developing neocortex. Sci. Rep. 6, 28998 (2016)(PMID:27383233)
- Mishima T et al. Determinants of effective lentivirus-driven microRNA expression in vivo. Sci Rep. 6, 33345 (2016)(PMID:27627961)
- Zaman MM et al. Arid5a exacerbates IFN-γ-mediated septic shock by stabilizing T-bet mRNA. PNAS. 113, 11543-11548 (2016)(PMID:27671645)
- Galipon J et al. Differential Binding of Three Major Human ADAR Isoforms to Coding and Long Non-Coding Transcripts. Genes (Basel). 8, E68 (2017)(PMID:28208661)
- Kotake Y et al. Y-box Binding Protein 1 Is Involved in Regulating the G2/M Phase of the Cell Cycle. Anticancer Res. 37, 1603-1608 (2017)(PMID:28373420)
- Wang ZN et al. High expression of PTBP1 promote invasion of colorectal cancer by alternative splicing of cortactin. Oncotarget 8, 36185-36202 (2017)(PMID:28404950)
- de Vasconcellos JF et al. IGF2BP1 overexpression causes fetal-like hemoglobin expression patterns in cultured human adult erythroblasts. PNAS. 114, E5664-E5672 (2017)(PMID:28652347)
- Lv J et al. Binding of LINE-1 RNA to PSF transcriptionally promotes GAGE6 and regulates cell proliferation and tumor formation in vitro. Exp Ther Med. 14, 1685-1691 (2017)(PMID:28810637)
- Brady LK et al. Chondroitin Sulfate Is Required for Onset and Offset of Critical Period Plasticity in Visual Cortex. PLoS Biol. 15, e2002623 (2017)(PMID:28961236)
- Hou X et al. Transcriptome analysis of hypoxic cancer cells uncovers intron retention in EIF2B5 as a mechanism to inhibit translation. Sci Rep. 7, 12646 (2017)(PMID:28974755)
- Yamaguchi T et al. The CCR4-NOT deadenylase complex controls Atg7-dependent cell death and heart function. Sci Signal. 11, eaan3638 (2018)(PMID:29438013)
- Fujita Y et al. KH-type splicing regulatory protein is involved in esophageal squamous cell carcinoma progression. Oncotarget 8, 101130-101145 (2017)(PMID:29254151)
- Cui J, Placzek WJ. PTBP1 enhances miR-101-guided AGO2 targeting to MCL1 and promotes miR-101-induced apoptosis. Cell Death Dis. 9, 552 (2018)(PMID:29748555)
- Sekiba K et al. DHX9 regulates production of hepatitis B virus-derived circular RNA and viral protein levels. Oncotarget. 9, 20953-20964 (2018)(PMID:29765512)
- Gu W et al. Dnd1-mediated epigenetic control of teratoma formation in mouse. Biol Open 7, bio032318 (2018)(PMID:29378702)
- Zhou A et al. RNA Binding Protein, HuR, Regulates SCN5A Expression Through Stabilizing MEF2C transcription factor mRNA. J Am Heart Assoc. 7, e007802 (2018)(PMID:29678826)
- Tajirika T et al. DEAD-Box Protein RNA-Helicase DDX6 Regulates the Expression of HER2 and FGFR2 at the Post-Transcriptional Step in Gastric Cancer Cells. Int J Mol Sci. 19, E2005 (2018)(PMID:29987267)
- Kuranaga Y et al. SRSF3, a Splicer of the PKM Gene, Regulates Cell Growth and Maintenance of Cancer-Specific Energy Metabolism in Colon Cancer Cells. Int J Mol Sci. 19, E3012 (2018)(PMID:30279379)
- Ishiguro S et al. Base-pairing probability in the microRNA stem region affects the binding and editing specificity of human A-to-I editing enzymes ADAR1-p110 and ADAR2. RNA Biol. 15, 976-989 (2018)(PMID:29950133)
- Hitachi K et al. Myogenin promoter-associated lncRNA Myoparr is essential for myogenic differentiation. EMBO Rep. (2019) In press.
(PMID:30622218)
- Tsujino T et al. MicroRNA-143/Musashi-2/KRAS cascade contributes positively to carcinogenesis in human bladder cancer. Cancer Sci. 110, 2189-2199 (2019)(PMID:31066120)
- Hu Y et al. The antibiotic clofoctol suppresses glioma stem cell proliferation by activating KLF13. J Clin Invest. 129, 3072-3085 (2019)(PMID:31112526)
- Araki K et al. TDP-43 regulates early-phase insulin secretion via CaV1.2-mediated exocytosis in islets. J Clin Invest. 130, 3578-3593 (2019)(PMID:31355778)
- Ma H et al. Noncoding RNA 886 alleviates tumor cellular immunological rejection in host C57BL/C mice. Cancer Med. 9, 5258-5271 (2020)(PMID:32476259)
- Senoo M et al. RNA-binding protein Ptbp1 regulates alternative splicing and transcriptome in spermatogonia and maintains spermatogenesis in concert with Nanos3. J Reprod Dev. 66, 459-467 (2020)(PMID:32624547)
|






 Citations
Citations Data Sheet
Data Sheet