| Data |

Properties of Fluorescent protein "hAG"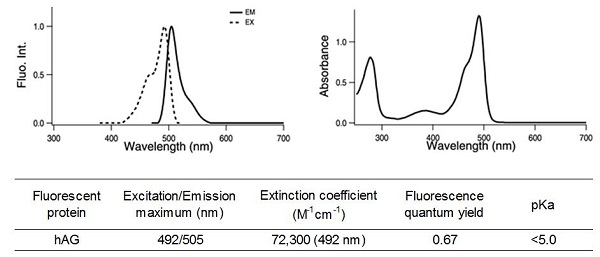

Plasmid Maps

Experimental example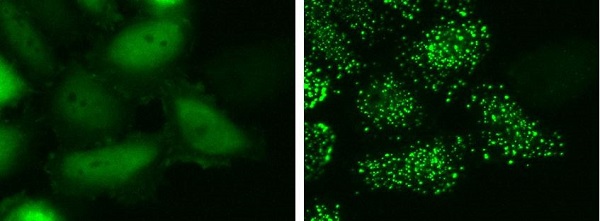
HeLa-S3 cells transiently expressing both Ash/FKBP12 and hAG/mTOR (FRB) were observed at 0 minute (left) and 12 minutes (right) after addition of 500 nM Rapamycic.
|
| Components |
pAsh/FKBP12 Plasmid, phAG/mTOR Plasmid |
| Intended use |
Fluoppi : Ash-hAG [mTOR-FKBP12] contains two plasmids for detecting mTOR(FRB)-Rapamycin-FKBP12 ternary complex in living cells. |
| Storage temp. |
-20°C |
Manufacturer |
MBL |
| Related products |
AM-8011M Fluoppi Ver.2 : Ash-hAG (Ash-MNL/MCL + hAG-MNL/MCL)
AM-VS0801M humanized Azami-Green for Fluoppi (phAG-MNL/MCL)
AM-VS0802M Monti-Red for Fluoppi (pMonti-Red-MNL/MCL)
AM-8201M Fluoppi : Ash-hAG [ p53-MDM2 ]
AM-8012M Fluoppi Ver.2 : Ash-Red (Ash-MNL/MCL + Monti-Red-MNL/MCL)
M223-3 Anti-Ash-tag mAb
AM-P0001 PPIs Detection Reagent : Fluoppi [ p53-MDM4 ]
AM-P0002 PPIs Detection Reagent : Fluoppi [ BclxL-BAK ]
AM-P0003 PPIs Detection Reagent : Fluoppi [ BclxL-BAX ]
AM-P0004 PPIs Detection Reagent : Fluoppi [ Bcl2-BAK ]
AM-P0005 PPIs Detection Reagent : Fluoppi [ Bcl2-BAX ]
AM-P0006 PPIs Detection Reagent : Fluoppi [ Mcl1-BAK ]
AM-P0007 PPIs Detection Reagent : Fluoppi [ Mcl1-BAX ]
AM-P0010 PPIs Detection Reagent : Fluoppi [ p50-p65 ]
AM-P0013 PPIs Detection Reagent : Fluoppi [ CDK4-Cyclin D1 ]
AM-P0014 PPIs Detection Reagent : Fluoppi [ CDK5-p25 ]
AM-P0015 PPIs Detection Reagent : Fluoppi [ JNK-JIP ]
AM-P0016 PPIs Detection Reagent : Fluoppi [ p21-PCNA ]
AM-P1002 PPIs Detection Reagent : Fluoppi [ BclxL-BAK ]
AM-P1003 PPIs Detection Reagent : Fluoppi [ BclxL-BAX ]
AM-P1004 PPIs Detection Reagent : Fluoppi [ Bcl2-BAK ]
AM-P1005 PPIs Detection Reagent : Fluoppi [ Bcl2-BAX ]
AM-P1006 PPIs Detection Reagent : Fluoppi [ Mcl1-BAK ]
AM-P1007 PPIs Detection Reagent : Fluoppi [ Mcl1-BAX ]
|
| Notes |
"Please note that the product will be shipped after we receive a completed and signed License Acknowledgement. Prior to shipping, our staff may contact you to verify the intended use of the product.
DNA sequence:
CoralHue® vectors full DNA sequence" |
| Citations |
- Koyano F et al. Ubiqutitin is phosphorylated by PINK1 to activate perkin. Nature. 510, 162-6 (2014)(PMID:24784582)
- Yamano K et al. Site-specific Interaction Mapping of Phosphorylated Ubiquitin to Uncover Parkin Activation. J Biol Chem. 290, 25199-211 (2015)(PMID:26260794)
- Asamitsu K et al. Quantification of the HIV transcriptional activator complex in live cells by image-based protein-protein interaction analysis. Genes Cells. 21, 706-16 (2016)(PMID:27193293)
- Watanabe T et al. Genetic visualization of protein interactions harnessing liquid phase transitions. Sci Rep. 7, 46380 (2017)(PMID:28406179)
|
| References |
- Choi J et al. Structure of the FKBP12-rapamycin complex interacting with the binding domain of human FRAP. Science 273, 239-42 (1996)(PMID: 8662507)
|
| Product category |
- Tools
- Fluorescent proteins
|





 Citations
Citations Data Sheet
Data Sheet
